Maintaining homeostasis requires exquisite control over the synthesis of new proteins, and accordingly cells have evolved several mechanisms that regulate the timing, levels, and localization of translation. In eukaryotic cells, these mechanisms include the use of 7-methylguanosine-capped (m7G) mRNAs and a cap-binding complex, eIF4F, that specifically recognizes this modification to drive initiation of protein synthesis. In contrast, viruses have long been thought to lack protein synthesis machinery, and instead co-opt host ribosomes through non-canonical initiation pathways and the use of unique mRNA features such as internal ribosome entry sites.
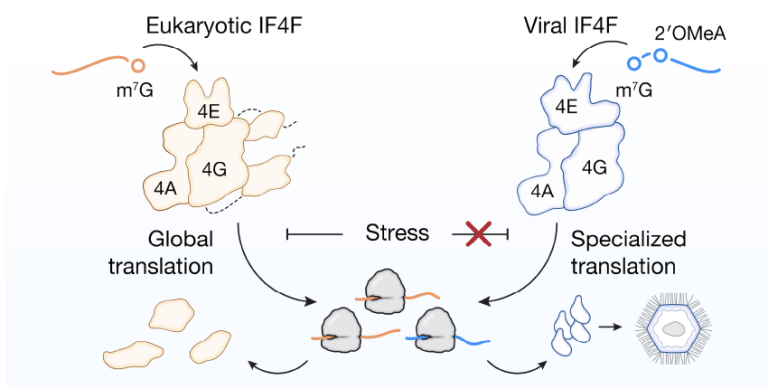
New work from the Lee lab upends this dogma by showing that so-called giant DNA viruses, which rival bacteria in terms of particle size and genomic content, encode a functional and essential viral cap-binding complex (vIF4F) that is homologous to eIF4F. During infection of the natural amoeba host, vIF4F associates with ribosomes to drive synthesis of the viral structural proteins late in the infection cycle. This temporal and transcript-specific regulation of translation is achieved partly through programmed recognition of viral mRNA cap structures, that beyond the canonical m7G modification, include a near-invariable 2′OMe-adenosine at the first position. Furthermore, giant DNA viruses that encode a cap-binding complex can maintain protein synthesis and viral replication under host stress conditions that would be highly detrimental to most mammalian viruses, suggesting a potential evolutionary role for the unusually complex form of translation regulation present in these viruses.
This work thus uncovers how horizontal transfer of core cellular genes to giant DNA viruses has led to the emergence of novel viral replication strategies that blur the boundaries of cellular and acellular biology. In essence, these cellular genes function as molecular “loanwords” that originally evolved within the regulatory and structural grammar of eukaryotic cells but have here been repurposed in a radically different context, revealing new rules that govern the language of life.