Most eukaryotic proteins are degraded by the 26S proteasome after modification with a polyubiquitin chain. Substrates lacking unstructured segments cannot be degraded directly and require prior unfolding by the Cdc48 ATPase (p97 or VCP in mammals) in complex with its ubiquitin-binding partner Ufd1-Npl4 (UN).
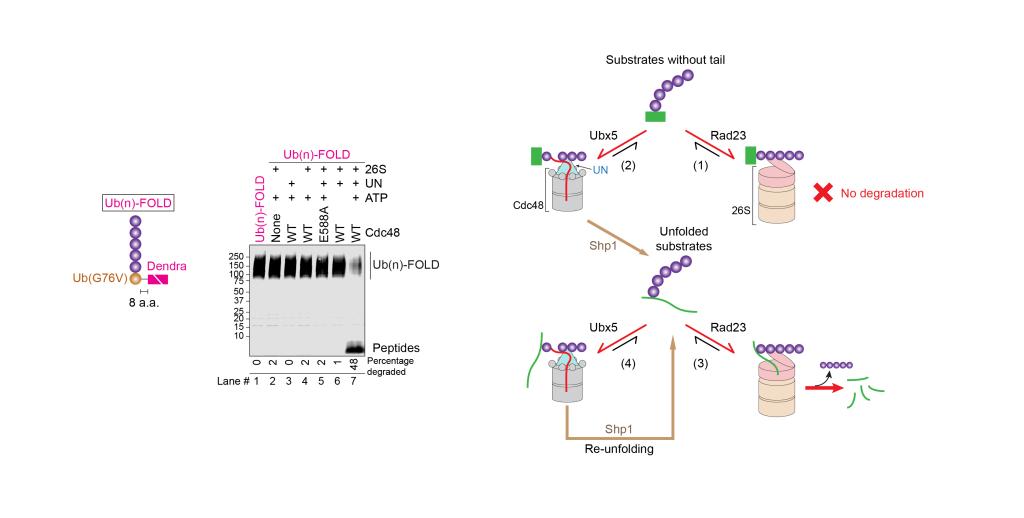
In this study, Hao Li, Zhejian Ji, Joao A. Paulo, Steven P. Gygi, and Tom A. Rapoport purified yeast components to reconstitute Cdc48-dependent degradation of well-folded model substrates by the proteasome. They show that a minimal system consists of the 26S proteasome, the Cdc48-UN ATPase complex, the proteasome cofactor Rad23, and the Cdc48 cofactors Ubx5 and Shp1. Rad23 and Ubx5 stimulate polyubiquitin binding to the 26S proteasome and the Cdc48-UN complex, respectively, allowing these machines to compete for substrates before and after their unfolding. Shp1 stimulates protein unfolding by the Cdc48-UN complex rather than substrate recruitment. Experiments in yeast cells confirm that many proteins undergo bidirectional substrate shuttling between the 26S proteasome and Cdc48 ATPase before being degraded.