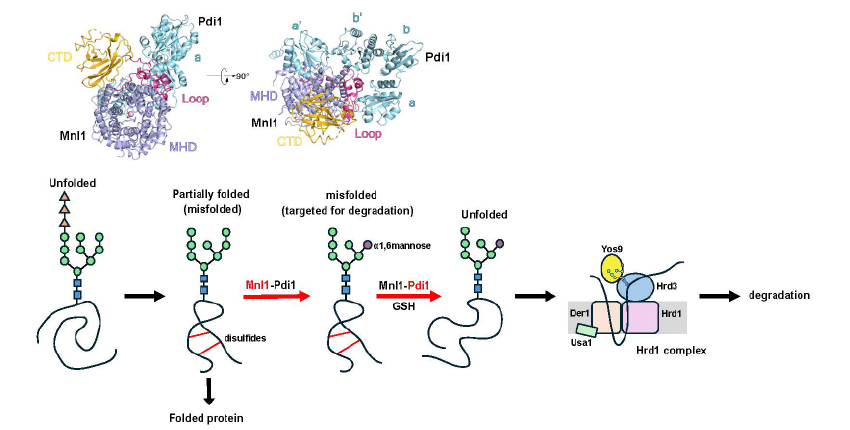
Misfolded glycoproteins in the endoplasmic reticulum (ER) lumen are translocated into the cytosol and degraded by the proteasome, a conserved process called ER-associated protein degradation (ERAD). The misfolded proteins are committed to degradation by a luminal mannosidase, called Mnl1 or Htm1 in Saccharomyces cerevisiae. Mnl1 generates a terminal a1,6-mannose residue in the glycan chain, which targets the glycoprotein to the Hrd1 complex for retro-translocation into the cytosol. Interestingly, Mnl1 is associated with protein disulfide isomerase (Pdi1), the enzyme that generates and isomerizes disulfide bonds. Zhao et al. used cryo-electron microscopy, biochemical and in vivo experiments to demonstrate that the Mnl1-Pdi1 complex first demannosylates misfolded, globular proteins that are recognized through the C-terminal domain (CTD) of Mnl1. Pdi1 causes the CTD to ignore completely unfolded polypeptides that have not yet undergone folding attempts. The disulfides of the demannosylated globular proteins are then reduced by the Pdi1 component of the complex. Mnl1 blocks the canonical oxidative function of Pdi1, as it prevents the re-oxidation of Pdi1 by Ero1. However, Pdi1 can still function as a reductase, using reduced glutathione as a reductant. Thus, the Pdi1 in the Mnl1-Pdi1 complex is the elusive disulfide reductase in ERAD, which generates unfolded polypeptides that can then be translocated across the membrane into the cytosol.